Project: Neural surrogate modeling of biomolecular condensate morphology
Description
Mapping the phase behavior of biomolecular condensates across a multi-dimensional parameter space is a fundamental challenge in soft matter science, relevant to materials design and drug delivery. Automated platforms can navigate this space using active machine learning, but current approaches rely on a binary acquisition signal, whether phase separation occurs or not, that discards the rich three-dimensional morphological information in each confocal image stack and cannot transfer knowledge across molecular variants.
This project aims to train a conditional generative model that predicts the full distribution over 3D condensate morphologies. We observe multi-modal behavior: identical input conditions can produce qualitatively distinct morphological outcomes that deterministic models cannot represent [1]. Flow matching, recently demonstrated for biological fluorescence image generation [2] and for physical systems with multiple coexisting solutions [3], provides the correct probabilistic framework for this problem.
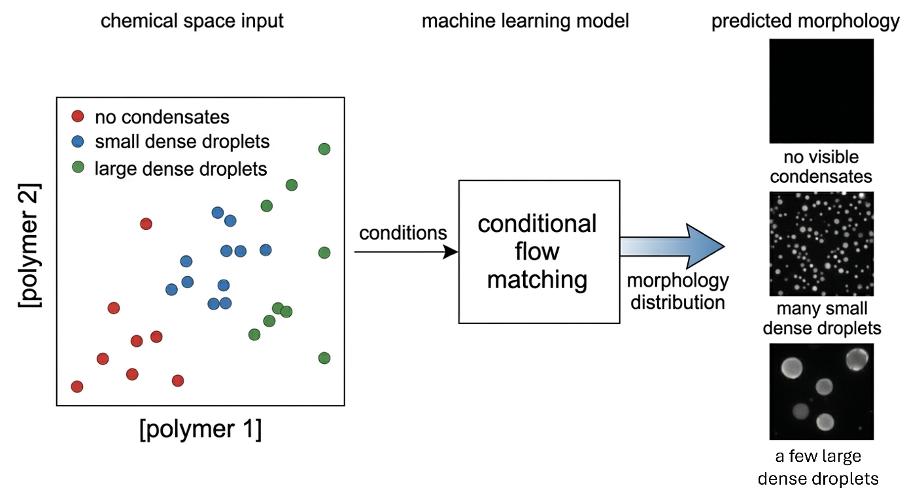
The model operates in a latent space learned by a 3D variational autoencoder compressing each confocal z-stack (36 slices, 256 × 256 pixels) into a compact morphological representation. A conditional flow matching model then maps experimental conditions to distributions over these latent vectors, trained jointly across nine molecular systems to enable generalization to variants not present in the training data. The student will benchmark the surrogate model on 32,400 z-stacks (already available), and integrate morphological uncertainty into an active learning acquisition function, comparing performance against the binary baseline used by current platforms.
[1] Hendriks, F., Rokoš, O., Doškář, M., Geers, M. G., & Menkovski, V. (2025). Equivariant Flow Matching for Symmetry-Breaking Bifurcation Problems. arXiv preprint arXiv:2509.03340.
[2] Konshin, N., Minartz, K., de Boer, J., & Menkovski, V. (2025). Micellangelo: a Generative Model of Cell-Topography Interactions. bioRxiv, 2025-11.
[3] Leurs, Yannick HA, Willem Van Den Hout, Andrea Gardin, Joost LJ Van Dongen, Andoni Rodriguez-Abetxuko, Nadia A. Erkamp, Jan CM van Hest, Francesca Grisoni, and Luc Brunsveld. "Automated navigation of condensate phase behavior with active machine learning." Nature Communications 16, no. 1 (2025): 9598.
Details
- Supervisor
-
 Vlado Menkovski
Vlado Menkovski
- Secondary supervisor
-
NENadia Erkamp
- Interested?
- Get in contact